On 19–21 August 2023, the Monash Centre for Electron Microscopy (MCEM) hosted the Electron Crystallography School 2023 (ECS2023) as a satellite event to the 26th IUCr Congress. The School was organised by the IUCr Commission on Electron Crystallography, with Hongyi Xu, Tatiana Gorelik, Mauro Gemmi and Laure Bourgeois as co-chairs.
The ECS2023 focused on two broad topics that have seen considerable development in the last decade or so: 4D scanning transmission electron microscopy (4D-STEM) and 3D/micro-electron diffraction (3D-ED). These developments have led to major advances in the characterisation of the atomic-scale structure of solid matter, from energy materials and green chemicals to glasses, proteins and pharmaceutical compounds.
The School was aimed at postgraduate students, early-career researchers and newcomers to the field. It offered a series of lectures on the fundamental theory underpinning 4D-STEM and 3D-ED, together with historical background, software demonstrations and hands-on tutorials. The intensive program also included a live demonstration on MCEM’s latest transmission electron microscope acquisition, the Thermo Fisher Scientific Spectra-φ. A session was dedicated to presentations from industry scientists. There was also a lively poster session and a tour of MCEM facilities.
ECS2023 included presentations from many distinguished international scientists including Professor Lukáš Palatinus from the Czech Academy of Sciences, shown here.
The School was very fortunate to have 20 distinguished scientists from both overseas and Australia as its speakers. Amongst them were Ute Kolb (Johannes Gutenberg Universität Mainz) and Jian-Min Zuo (University of Illinois), past and present recipients of the Gjønnes Medal in Electron Crystallography, the highest distinction in the field.
Over 70 delegates from 13 countries and 4 continents attended the School, with students and researchers from countries as far away as Brazil, Belgium and China. This included a significant cohort from the X-ray crystallography community. Feedback from attendees was very positive, with appreciation shown not only for the quality of the program but also for the opportunity to meet with the broad crystallography community.
The organising committee is grateful for the generous support from the following sponsors: the IUCr, Microscopy Australia, the Society of Crystallographers in Australia and New Zealand (SCANZ), Rigaku, JEOL, Thermo Fisher Scientific, Dectris, ReadCrystal, Gatan Ametek and NanoMEGAS. The IUCr grants supported 13 PhD students and postdoctoral researchers, 4 of whom came from overseas (Belgium, Germany and China).
More detail can be found on the ECS2023 website.
Reports from IUCr grant recipients
Loreibelle Abian, Monash University, Australia
This student recipient was given the opportunity to attend the Electron Crystallography School 2023 last 19–21 August 2023 at Monash University-Clayton Campus, Clayton, Victoria, Australia. The said endeavour was attended by postgraduate students and early-career researchers coming from different countries as well as remarkable presenters from diverse fields of electron crystallography. The invited speakers talked about the advances in electron crystallography and cutting-edge technologies behind the state-of-the art instruments used in electron microscopy. The 3-day activity enabled the student to widen her horizon in the area of electron diffraction as she learned about other electron microscopy methods aside from the convergent-beam electron diffraction (CBED) technique which she currently employs in her research in investigating chemical bonding in inhomogeneous materials. The discussions during the lectures tackled CBED, scanning electron nanodiffraction (SEND), 4D scanning transmission electron microscopy (4D-STEM), 3D electron diffraction (3D-ED) and micro-electron diffraction (MicroED). The presentations were excellently delivered in that they undertook not only the fundamentals in crystallography and microscopy but also the latest research into which a speaker’s group is involved. Moreover, the participants were provided with a platform to share their research during the poster presentation on the evening of the first day where they were able to acquire insights and recommendations from renowned scientists and experts who graced the said event. The student would like to extend her gratitude for the sponsorship to join ECS which has given her an avenue to learn more about her research and to connect and build networks with fellow researchers.
Bikas Aryal, Monash University, Australia
The Electron Crystallography School (ECS) held at Monash University provided the attendees with both theoretical concepts of electron diffraction through excellent lectures and practical concepts through live software and TEM operation demonstrations. The ECS covered a wide range of topics related to electron diffraction and crystallography, such as selected area electron diffraction (SAED), scanning electron nanodiffraction (SEND), scanning transmission electron microscopy (STEM), quantitative convergent-beam electron diffraction (QCBED), MicroED, ptychography and 4D-STEM. By participating in ECS, I got a chance to understand the fundamentals of electron diffraction and advances in different techniques like 4D-STEM, SEND, MicroED and ptychography. As my research focuses on chemical bonding studies using QCBED, I enjoyed the talks “Direct observation of d-holes in Cu2O” and “Measuring vacancy concentrations, chemical bonding and lattice contraction around nanovoids in aluminium by QCBED”.
Furthermore, the ECS provided me with an opportunity to meet world-renowned experts such as Jian-Min Zuo, Randi Holmestad, Lukáš Palatinus, Ute Kolb and Colin Ophus. I discussed future collaborations with Professor Zuo. I also met many early-career researchers and representatives from different companies. We discussed new research directions and explored industrial collaborations.
I am grateful for the International Union of Crystallography's (IUCr) financial assistance, which greatly aided in covering my travel costs. The main benefit of these scholarships is their capacity to open up educational opportunities and scientific events to a wide range of participants.
Wei Chao, Monash University, Australia
I am so honoured and grateful for being selected as a recipient of an IUCr grant, which has allowed me to participate in the Electron Crystallography School. The school seamlessly integrated applications of electron diffraction in both materials science and biology, providing me with a highly educational and inspirational experience. This unique opportunity has enabled me to engage with presentations by pioneers in convergent-beam electron diffraction, 3D-ED/MicroED, offering invaluable insights into the beauty of diffraction across diverse disciplines.
The content of the school was so well organized. As a researcher in materials science, I not only enjoyed outstanding talks within my field but also had the chance to explore a new area of study. Despite my limited knowledge of MicroED, I learned and enjoyed a lot from the well-structured and comprehensive talks. I am sure that students and young researchers in the MicroED field would similarly find value in the discussions on convergent-beam electron diffraction.
The inclusion of hands-on workshops, focused on software packages, added a practical dimension that I found particularly enriching.
I would like to express my sincere gratitude to the organizers for their dedicated efforts in making this event a success. It is such a valuable experience in my career. I also wish to express my deep appreciation to the committee for the grant, which makes my attendance at this wonderful event possible.
Mostafa Eid Abdelmaboud Ahmed Ali, Monash University, Australia
The Electron Crystallography School (ECS), held between 19 and 21 August 2023 at Monash University, aimed to expand our knowledge of the electron crystallography area and provided us with a unique platform to explore it. The ECS covered a wide range of topics, providing attendees with both theoretical information through excellent lectures and practical skills through software demonstrations. For example, the lectures covered fundamental principles of electron crystallography techniques such as selected area electron diffraction (SAED), scanning electron nanodiffraction (SEND), scanning transmission electron microscopy (STEM), convergent-beam electron diffraction (CBED), quantitative convergent-beam electron diffraction (QCBED) and 4D-STEM.
Furthermore, the lectures and hands-on sessions (software demos) were presented by renowned experts and academics from top universities around the world, ensuring a thorough understanding of the subject matter. For example, Professor Jian-Min Zuo, a pioneer and brilliant in the field of electron crystallography, was among those present. I met him at the school, introduced myself and my proposal to him, and discussed some electron diffraction topics with him. This was very much appreciated.
My PhD research focuses on CBED and QCBED approaches, therefore I really learned a lot about the current issues and applications surrounding these methods, as well as the difficulties in gathering and interpreting electron diffraction data and ways to get over these obstacles. For instance, as one of the QCBED applications, I was interested in Professor Zuo's lecture “Direct observation of d-holes in Cu2O” and Professor Philip Nakashima's session “Vacancy mapping in Al structure”.
Besides the mentioned benefits of attending ECS, I was very happy for the International Union of Crystallography (IUCr) financial support, which helped me a lot to cover travel expenses. The primary impact of such grants lies in their ability to make educational opportunities and scientific events accessible to a diverse range of participants.
Weilun Li, Monash University, Australia
Overview
The Electron Crystallography School proved to be an enriching experience, offering a comprehensive understanding of advanced techniques in structural analysis using electron crystallography. Held at Monash, Melbourne, the school gathered a diverse group of researchers, scientists and students eager to delve into the intricacies of this cutting-edge field.
Programs and lectures
The school commenced with insightful lectures by eminent speakers who shared their expertise on the theoretical foundations of electron crystallography, its applications and recent advancements in the field. These lectures provided a solid foundation for participants, irrespective of their prior knowledge, ensuring that everyone could grasp the complexities of electron crystallography. The combination of CBED and 4D-STEM on the first day and 3D-ED and MicroED on the following day proved to be extremely effective.
Hands-on sessions
A highlight of the Electron Crystallography School was the demo lab sessions on the state-of-the-art TEM at Monash Centre for Electron Microscopy, Spectra-Phi, as well as data processing sessions on 4D-STEM and MicroED. Participants were guided through practical exercises using electron microscopes and image processing software. This hands-on experience was invaluable, allowing us to apply theoretical knowledge to real-world scenarios. The instructors were patient and skilled, providing personalized guidance to ensure that each participant could make the most of the practical sessions.
Meal/tea breaks
The school fostered a collaborative and engaging environment, facilitating interactions among participants, instructors and guest speakers. Networking sessions and group activities allowed attendees to exchange ideas, discuss challenges and explore potential collaborations. The diversity of backgrounds among participants enhanced the overall learning experience, providing a holistic perspective on the applications and challenges of electron crystallography in various scientific disciplines.
Poster sessions
The Electron Crystallography School also provided a platform for students and early-career researchers to showcase their research projects and findings. The poster sessions enabled attendees to gain deeper insights into the ongoing research in the field. This exposure to diverse projects and methodologies broadened our understanding of the potential applications of electron crystallography across different scientific domains.
Conclusions
The Electron Crystallography School was a comprehensive and well-organized program that delivered a profound learning experience. The combination of theoretical lectures, hands-on sessions, networking opportunities and exposure to cutting-edge research positioned participants to apply their new-found knowledge and skills in their respective fields. The school not only expanded our understanding of electron crystallography but also cultivated a sense of community among participants, fostering future collaborations and advancements in this dynamic field. I am confident that the knowledge gained at the Electron Crystallography School will significantly contribute to my research and academic endeavours.
Huyen Pham, Monash University, Australia
My name is Huyen Pham, a research fellow at the School of Physics and Astronomy, Monash University, Australia. I am extremely grateful and excited to have been part of the community attending the Electron Crystallography School, particularly after the hard period of COVID-19 lockdown and travel restriction. As a dedicated researcher in the field of materials characterisation by electron microscopy, I am always hungry to approach and acquire more knowledge in this topic. Participating in this event gave me an opportunity not only to expand my personal network but also to learn more fundamental knowledge and catch up with up-to-date progress in electron crystallography from famous scientists around the world. The school was well organized with a series of lectures on a wide range of topics including the fundamental theory of convergent-beam electron diffraction (CBED), 4D scanning transmission electron microscopy (4D-STEM) and 3D electron diffraction (3D-ED) together with historical background, software demonstrations and hands-on tutorials. This experience will truly support my growth and passion to be a researcher in electron microscopy. I also would like to thank the IUCr for committing to give me an IUCr grant, which strongly supports my travel costs and other necessary expenses.
Sepideh Rahimi, University of Antwerp, Belgium
I'm writing to convey my heartfelt appreciation for the Young and Early Career Scientist Award I had the privilege of receiving from the IUCr during the Electron Crystallography School at Monash University. This recognition has played a crucial role in facilitating my participation in this educational program. Your support means a lot to me, and I'm sincerely grateful for it. The experiences I gained have been immensely enriching and have contributed to my professional and personal growth.
The school allowed me to deepen my knowledge of 4D-STEM and CBED, gave me insights about developing my in-situ liquid experiments and provided valuable comments and ideas from famous experts in the field. I also had the opportunity to establish connections and collaborate with peers and mentors from around the world.
In terms of personal development, the experience broadened my horizons and enhanced my intercultural skills. It was an unforgettable experience that will undoubtedly benefit my future endeavors.
I am profoundly grateful for the support the Young and Early Career Scientist Award provided, which made this transformative journey possible. I am committed to utilizing the knowledge and experiences gained during this trip to further advance my research and contribute to the scientific community.
Once again, thank you for believing in my potential and supporting my academic and professional pursuits. I am truly honored to be a recipient of this prestigious award.
Please convey my gratitude to the members of the selection committee and all those who make these awards possible. I look forward to the opportunity to share the outcomes of this experience with you shortly.
Alaa Shaikhqasem, Martin-Luther Universität Halle-Wittenberg, Germany
I had the privilege to attend the Electron Crystallography School (ECS2023) which took place in August 2023 at the prestigious Monash University. My primary focus lies in 3D electron diffraction. The school presented a distinctive opportunity to connect with pioneering scientists who are actively shaping and sharing cutting-edge advancements in this field. Furthermore, this international event brought together participants from diverse backgrounds and experiences, creating a rich environment for exchanging ideas and establishing collaborations. The ECS2023 school provided me with an opportunity to present my work on 3D electron diffraction of protein crystals. Additionally, throughout the program, I acquired an extensive knowledge base in the field of electron diffraction, covering a wide range of topics including the following:
- Advances in methods development and application
- Sample preparation
- 3D electron diffraction data collection
- Data processing for small molecules and macromolecules
- Structure refinement and model validation.
How did the IUCr grant help?
Traveling from Germany posed financial challenges due to limited funds for travel, accommodation and expenses during my stay for the school. In this context, the IUCr grant proved to be a significant source of support, facilitating attendance and accommodation during the school.
I feel incredibly privileged to have been selected for the IUCr award, and I am grateful for the opportunity. I would like to express my heartfelt appreciation to Professor Laure Bourgeois and Dr Hongyi Xu for organizing the school and for their exceptional support and guidance throughout the duration of the event.
Yu Zhang, Monash University, Australia
As a student specialising in electron crystallography, receiving the IUCr Young and Early Career Scientist Award to attend the Electron Crystallography School 2023 has been a truly transformative experience. It provided me with a unique platform to engage with preeminent experts in the field, leading to enriching discussions and invaluable learning opportunities.
The sessions were nothing short of illuminating, particularly in revealing the remarkable capabilities of 4D-STEM and MicroED/3D-ED techniques. I was captivated by the hands-on laboratory demonstrations of 4D-STEM data collection led by Matthew Weyland, as well as the insightful discussions on post-processing of 4D-STEM by Jian-Min Zuo and Bryan Esser, each employing different software platforms, showcasing the diverse approaches to 4D-STEM data processing.
Moreover, the demonstrations by Colin Ophus, Scott Findlay and Hamish Brown, demonstrating the simulation of 4D-STEM using Python, were exceptionally intriguing. The talks covering precession electron diffraction and MicroED/3D-ED further deepened my understanding of these cutting-edge topics, highlighting the transformative potential of these techniques in structure determination, and significantly enhancing my insights. The subsequent industry speeches provided me with a profound understanding of the current priorities of leading companies, which has inspired my future learning and research. In summary, this masterclass has offered me a broader perspective on electron crystallography, allowing me to venture beyond the confines of my own research domain. I am sincerely grateful to IUCr for affording me this remarkable opportunity to participate in ECS2023.
Cheng Zhao, Monash University, Australia
The electron crystallography school (ECS) was truly an unforgettable experience. It was the first time that I had met with academics and researchers that I had repeatedly read about in the literature and textbooks.
ECS was well organised from the very beginning to the end. The program included a variety of areas in electron microscopy/electron diffraction. As an electron microscopy student, I was amazed and inspired by the continuous exploration people had put into developing techniques and even building up their own equipment for certain experiments. I was also very interested in all the demonstrations on newly developed simulation techniques. The ‘new for me’ information on three-dimensional electron diffraction was also a very interesting aspect that I had never thought to be so useful before.
Another point that made this school so different was that it started with very fundamental knowledge with few information barriers that I’m sure helped a lot of students who were not very familiar with electron crystallography.
Some of the most inspiring mentality that I learned was to always have faith in yourself (research-wise) and not be discouraged by other people’s words. On the other hand, one should have an open heart to the work of others and try your best to appreciate their work rather than doubting it.
This grant was of great help in covering my travel expenses. I am sincerely thankful to the selection committee.
Yi Zhou, ShanghaiTech University, China
This was my first time in attending such an overseas international conference. I had a fantastic experience at ECS2023, because of not only the thrilling lectures on both fundamental and cutting-edge science and technology about convergent-beam electron diffraction (CBED), four-dimensional scanning electron microscopy (4D-STEM) and three-dimensional electron diffraction (3D-ED) but also the chance to discuss face to face with pioneers and talented young scientists from all around the world in these fields.
During the school, I was very impressed by the quantitative analysis of CBED data by Professor Jian-Min Zuo from the University of Illinois at Urbana-Champaign (UIUC) and Professor Philip Nakashima from Monash Centre for Electron Microscopy (MCEM). The unit cell, symmetry, sample thickness, atom positions and even orbitals can be analyzed. There were also a lot of lectures that showed us many useful software packages for processing 4D-STEM data with the speakers giving detailed instructions on how to use the software. Although I don’t work on this field, it still inspired me a lot. Lectures on 3D-ED and kinematical and dynamical structure refinement taught me a lot, which will benefit my experiments very much.
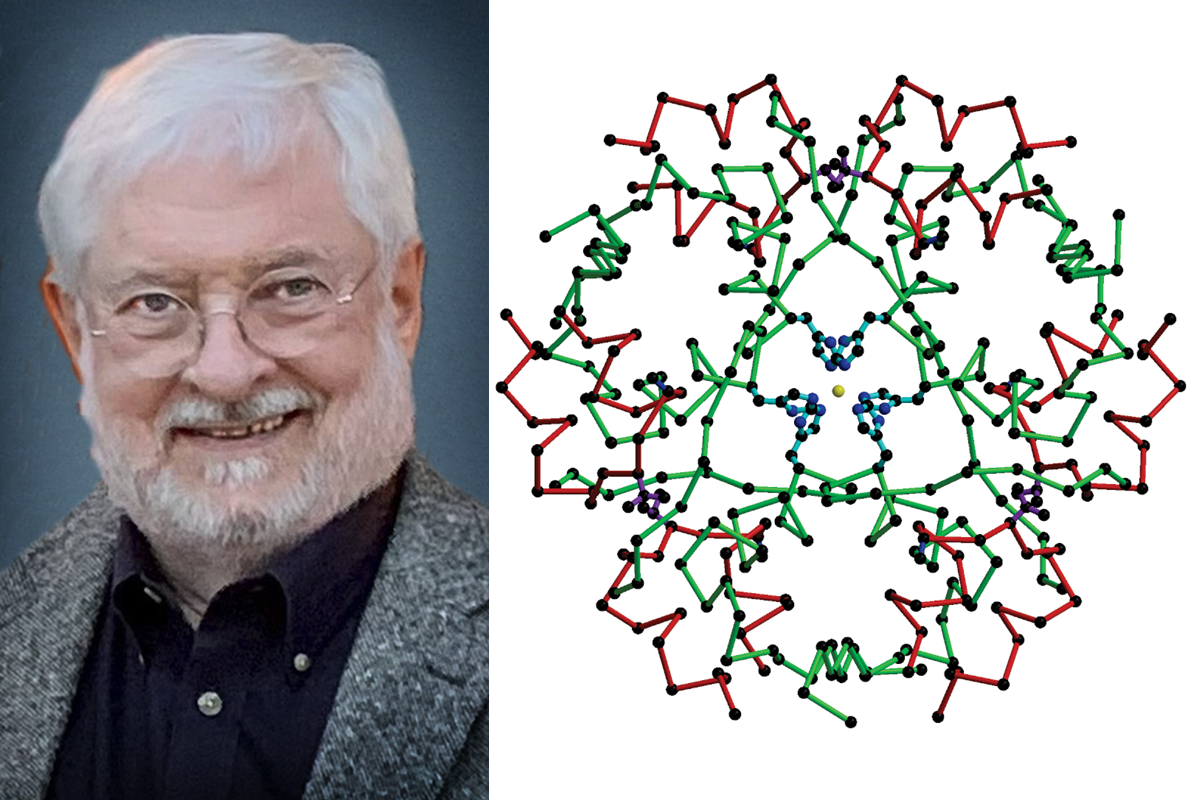
I was very honored to give a poster presentation on the evening of the first day of this electron crystallography school to present my up-to-date work about locating the guest species on metal–organic frameworks by 3D-ED. Several other talented young peers also presented their beautiful work, covering different materials and various techniques. It’s really good to know what others are doing and communicate with them.
Finally, I would like to say thanks to the organizers, Professor Laure Bourgeois and Hongyi Xu, for providing me with such a good platform to learn from others and promote myself. At the end of this school, I was awarded an “IUCr Young and Early Career Scientist Award”. This IUCr grant not only provided me with financial support but also gave me a lot of motivation. This is a kind of recognition for my previous research and encouragement for my future career.







![[IUCr2026 logo]](https://www.iucr.org/__data/assets/image/0005/158198/IUCr2026_Logo_RGB.jpeg)
![[AsCA logo]](https://www.iucr.org/__data/assets/image/0004/3955/asca.jpg)
![[ECA logo]](https://www.iucr.org/__data/assets/image/0005/3956/ecalogo.gif)
![[LACA_logo.jpg]](https://www.iucr.org/__data/assets/image/0016/111544/LACA_logo.jpg)











![[Fig. 1]](https://www.iucr.org/__data/assets/image/0016/158002/IUCr_report_ECS2023_presentation.png)

![[Fig. 1]](https://www.iucr.org/__data/assets/image/0008/157967/Fig.-1.png)
![[Fig. 1]](https://www.iucr.org/__data/assets/image/0009/157950/Fig.-1.jpg) In good voice. From left to right, Wilson Greatbatch, Herbert Hauptman, Bob Blessing and Lodovico Riva di Sanseverino sing at the 75th birthday celebration for Nobel laureates Herbert Hauptman and Jerome Karle in Buffalo in 1992. Photo contributed to the IUCr gallery by W. L. Duax.
In good voice. From left to right, Wilson Greatbatch, Herbert Hauptman, Bob Blessing and Lodovico Riva di Sanseverino sing at the 75th birthday celebration for Nobel laureates Herbert Hauptman and Jerome Karle in Buffalo in 1992. Photo contributed to the IUCr gallery by W. L. Duax.
![[Fig. 2]](https://www.iucr.org/__data/assets/image/0010/157951/Fig.-2.jpg) From left to right, Akihito Yamono, Tony Linden, Bob Blessing, Ludger Häming, Ton Spek, David Watkin and Larry Falvello, speakers at the Microsymposium on Absorption Corrections at the 18th IUCr Congress in Glasgow, UK, in 1999.
From left to right, Akihito Yamono, Tony Linden, Bob Blessing, Ludger Häming, Ton Spek, David Watkin and Larry Falvello, speakers at the Microsymposium on Absorption Corrections at the 18th IUCr Congress in Glasgow, UK, in 1999.